General information on SARS-CoV-2 RBD of Spike protein, K417N, L452R, T478K – AY.1/AY.2 lineage – Indian Delta plus variant
The Delta Plus variant was first identified after routine scanning of genomic variations in the lineage B.1.617.2 (original Delta variant) first reported in India. This new variant, which corresponds to lineages AY.1 and AY.2 (the first more abundant than the second), was shown to carry mutation K417N on its receptor-binding domain (RBD) in addition to mutations L452R and T478K previously reported on the original Delta variant.
The K417N amino acid substitution was previously reported in the Beta variant or lineage B.1.351 first identified in South Africa. This change occurs close to a domain targeted by neutralizing antibodies, for this reason, experts believe it might contribute to an enhanced immune evasion in SARS-CoV-2. This may have a synergistic effect in conjugation with mutation L452R, another substitution that is known to contribute to immune evasion in SARS-CoV-2. In contrast, the T478K mutation occurs within the interface between the RBD and the human receptor ACE2. Several reports indicate that the global prevalence of this amino acid change is slowly increasing hinting at its higher transmissibility and perhaps greater affinity towards the human receptor.
The prevalence of lineage AY.1/AY.2 has been steadily increasing in several European countries including the UK, Switzerland, Portugal, and France, and the United States. More recently, Asian countries including India have reported the first cases of the new Delta plus variant. Although previous reports of the Delta have indicated this variant to be more transmissible and likelier to cause a surge in hospitalizations, it remains to be seen if variant Delta plus will cause similar effects. Given the additional immune-escape-conferring mutation, lineage AY.1 might have a slightly better fitness advantage in comparison to the Delta variant. However, more studies are necessary to understand the concerted effect of the three mutations into SARS-CoV-2 transmissibility, virulence, and ability to evade the immune response generated by different COVID-19 vaccines.


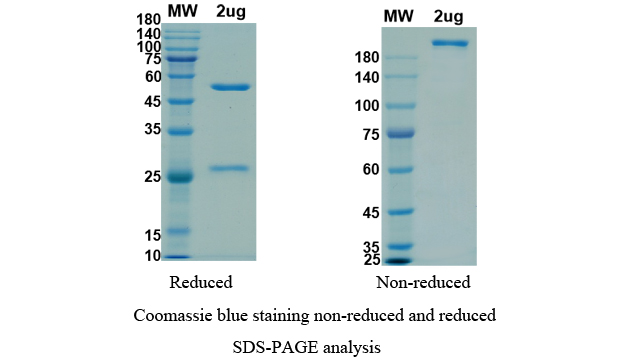
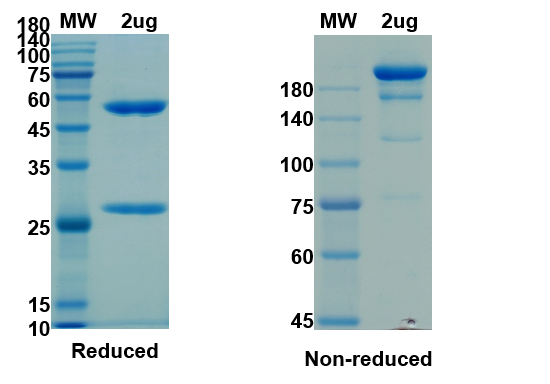
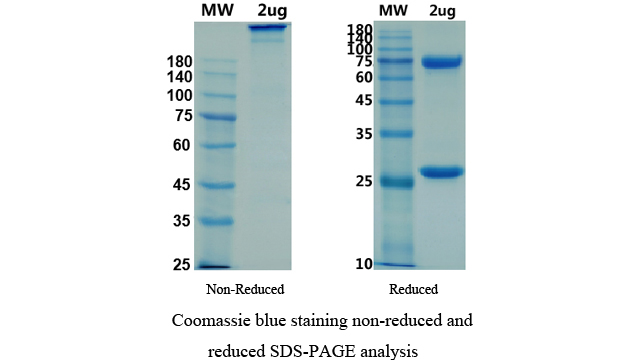
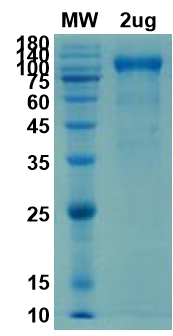
There are no reviews yet.