General information on RBD of VUI V1 UK variant (VOC-202012/01): N501Y
The N501Y mutation in the receptor-binding domain (RBD) has been detected in an emerging variant of SARS-CoV-2. The variant, known as the B.1.1.7 lineage, Variant of Concern (VOC) 202012/01, or clade 20B/501Y.V1, was first discovered in Kent, United Kingdom (UK), in mid-September 2020 and its frequency and distribution continued to grow since then.
The growing number of infections caused by this variant of the new coronavirus has raised causes for concern. The mutation N501Y is located in one of six key residues within the RBD and it has been linked to an enhanced binding affinity towards human ACE2 (angiotensin-converting enzyme 2). The enhancement can also explain the rapid expansion of its geographic distribution and potentially higher infectivity in comparison to other variants of the COVID-19 virus.
Aside from the N501Y mutation, the B.1.1.7 lineage has accumulated 16 additional non-synonymous mutations and deletions within the spike protein (but outside the 320-541 region). The most concerning ones besides the N501Y include:
- The 69-70del in the N terminal domain of the spike, which has been previously been linked with an enhanced ability to evade the human immune system
- The P681H mutation adjacent to the furin cleavage site of the spike, a location with high biological significance due to its role in spike cleavage and membrane fusion process
These cumulative mutations are hypothesized to contribute to the increased infectivity and rapid spread of the B.1.1.7 lineage. However, it is still unclear whether this large number of changes in the spike will impact disease severity or even hinder the efficiency of current COVID-19 vaccines.
The 69-70del has also been a cause of difficulty in effective RT-PCR detection kits that target the spike protein of SARS-CoV-2. This has created the need to develop a new assay that will allow the accurate surveillance of the B.1.1.7 variant. Moreover, further studies are needed to elucidate the impact of this set of mutations in the progression of the COVID-19 disease.


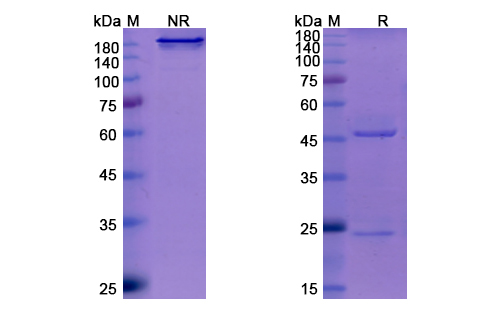
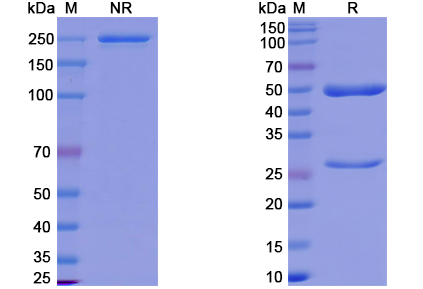
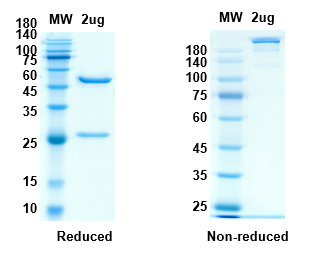
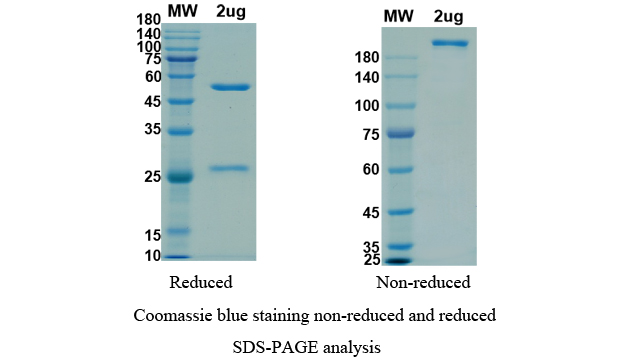
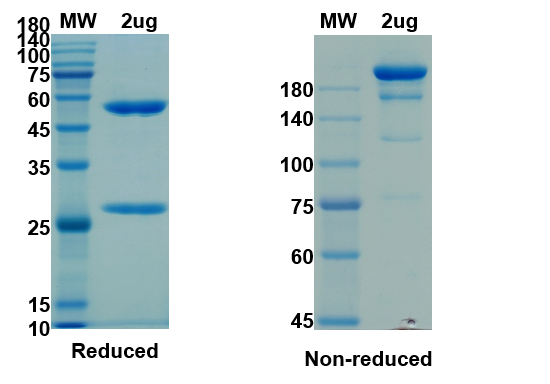
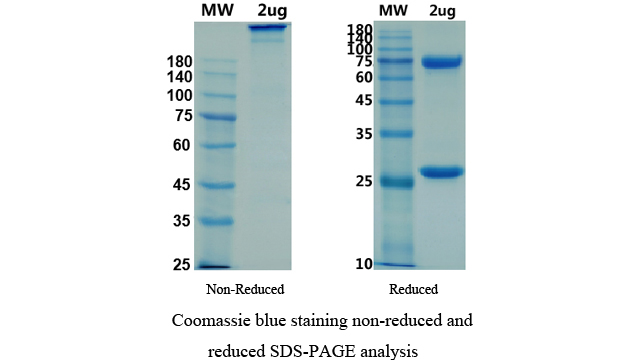
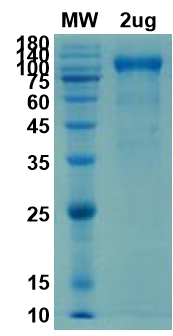
There are no reviews yet.