General information on SARS-CoV-2 RBD of Spike protein, K417N, E484K, N501Y
The accumulation of mutations K417N, E484K and N501Y in the receptor-binding domain (RBD) of the spike in the new coronavirus, SARS-CoV-2, has been recently described in an emerging variant of the virus. The variant, commonly known as the South African variant or the 501.V2 variant, may have emerged in early August 2020 in Nelson Mandela Bay, South Africa, according to spatiotemporal phylogeographic analysis. However, it was only identified as a Variant of Concern (VOC) by national authorities in mid-December 2020 and has since spread geographically across several provinces and countries.
Observational evidence provided by official authorities in South Africa indicates a potential shift in the clinical epidemiological scenario. Noticeably, clinicians observed a larger proportion of younger patients with no co-morbidities presenting critical illness. Moreover, recent data indicate that variant 501.V2 is the key responsible for causing a resurgence of COVID-19 infections and consequently driving the second wave of the pandemic in this region.
The N501Y mutation had been previously reported in another emerging variant of SARS-CoV-2 known as the B.1.1.7 lineage. This change within the receptor-binding motif (the main functional motif) is known to enhance the binding affinity of the virus to its natural human receptor – ACE2 (angiotensin-converting enzyme 2). ACE2 affinity is expected to be further enhanced by the presence of an additional non-synonymous mutation within this motif, the E484K mutation. This mutation was considered uncommon (less than 0.02% of the sequences outside South Africa) prior to the identification of the 501.V2 variant.
Further analysis of the third key mutation, mutation K417N, indicates this change is expected to have minimal impact on binding affinity to the human ACE2. However, its location is cause for concern. Prior studies indicate that the K417N mutation occurs within the area considered the main target of neutralizing antibodies (NAbs). For this reason, the mutation is expected to abolish interaction with certain NAbs and likely contribute to immune evasion.


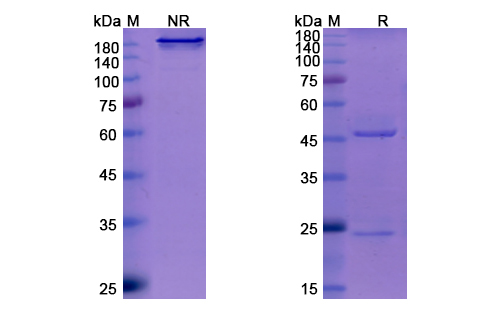
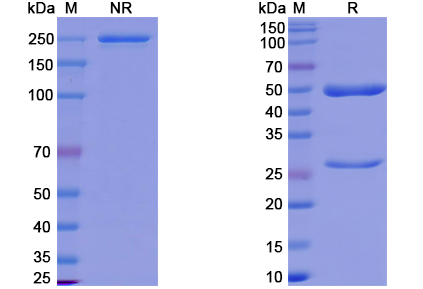
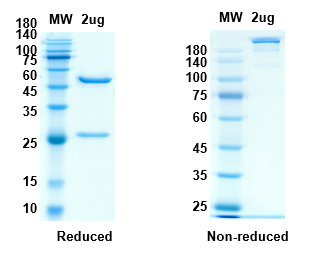
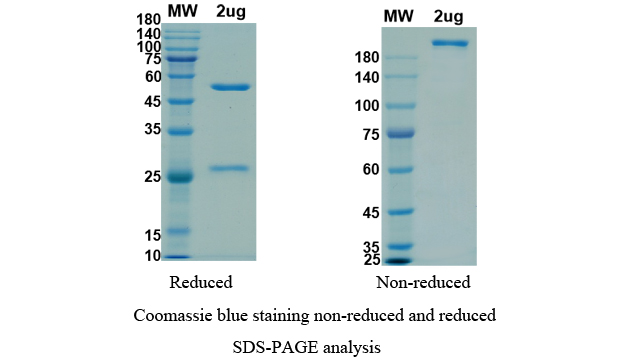
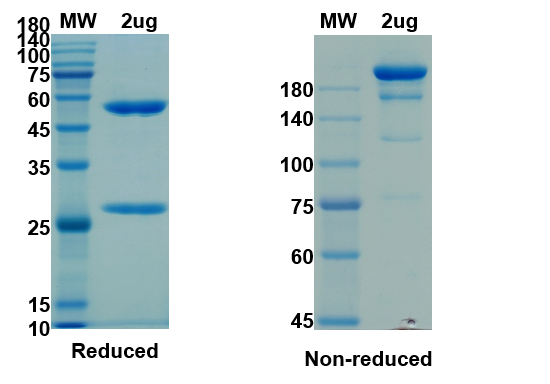
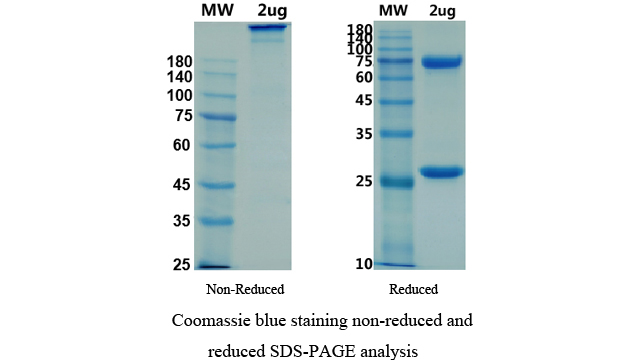
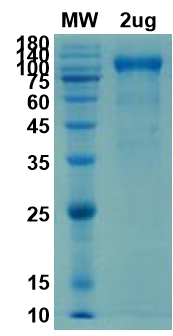
There are no reviews yet.