 Antibody production
Antibody production
In vivo vs. in vitro techniques for custom antibody generation
The first stage of custom antibody generation consists of the creation and maturation of these molecules in vivo (immunization) or in vitro (e.g. phage display). Each of these approaches to antibody production has different advantages and limitations. Understanding them is the key step of any successful project.
In vivo custom antibody generation
In vivo generation of antibodies requires immunization and subsequent harvesting of the host’s effector B cells or antiserum. When both the creation and the maturation of antibodies take place in vivo, it is possible to generate, in a cost-effective way, a number of different antibody molecules with high affinity to different epitopes of the same antigen (i.e. different parts of the antigen).
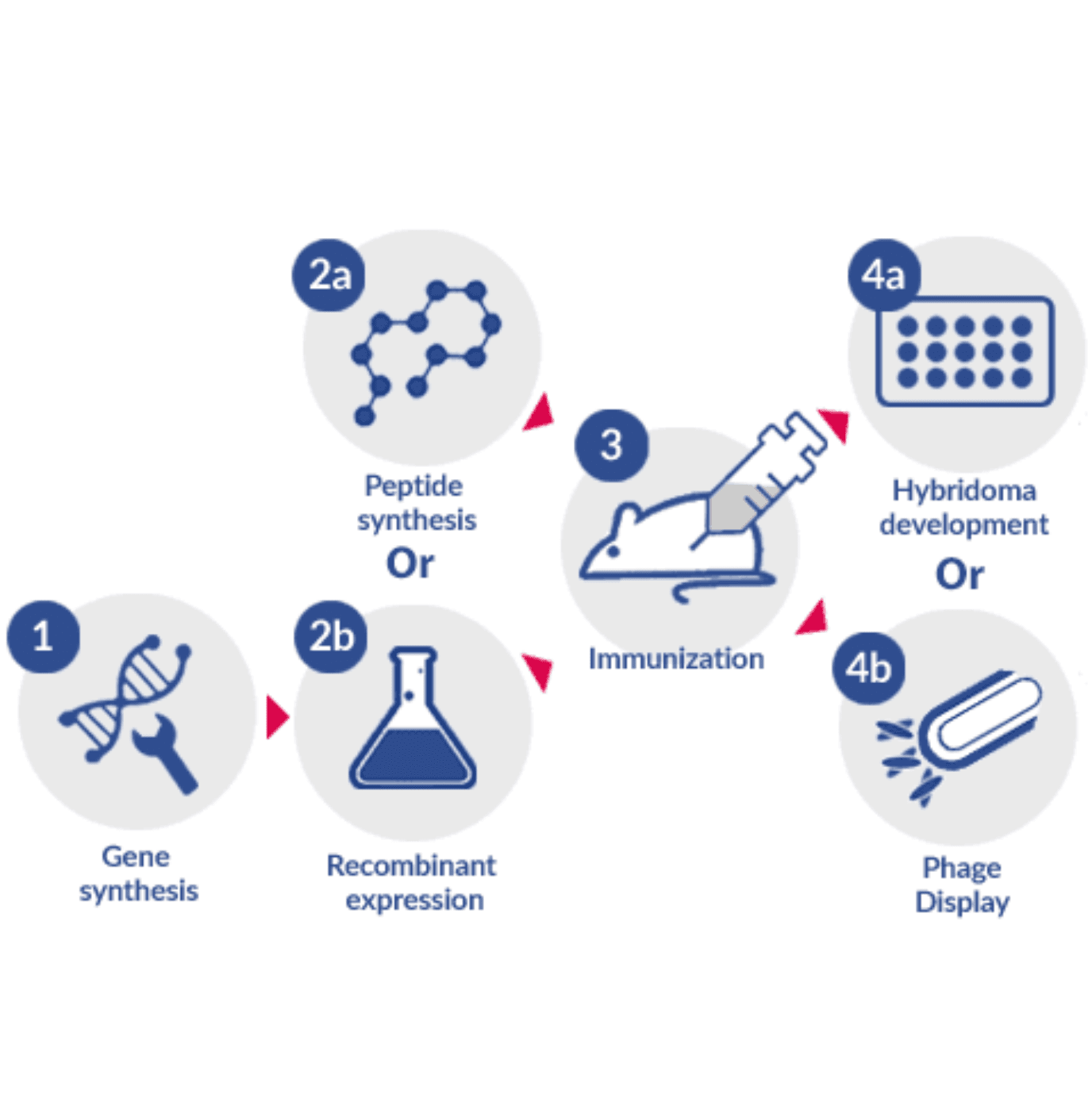
In this process, the clonal selection of the most effective B cell germlines is performed by the host’s immune system, and it results in the generation of several highly specific and mature germlines that can be harvested for further development.
But because the process of maturation takes place in vivo, it cannot be directly controlled. For this reason, the quality of in vivo generated antibodies relies heavily on the design of an effective immunization strategy. Therefore, the synthesis and purification of an adequate antigen, choice of an adequate antigen carrier, selection of an optimal adjuvant and planning of an effective immunization schedule are the most important steps of this process. And because several types of molecules can be used for immunization (e.g. proteins, peptides, DNA), some degree of control can be exerted at this stage by choosing which part of the antigen will be presented to the host’s immune system.
There are also important drawbacks in using animal hosts to generate antibodies. For instance, the use of non-immunogenic antigens or antigens that cause adverse effects on the host may fail to challenge their immune system and start an adequate immune response, which limits the type and structures of antigens that can be used for in vivo development.
At this point, if the purpose of the antibody generation is the development of biosensors for detection or as antitoxins, it is sufficient to harvest and purify antiserum of the immunized host. This procedure generates a mixture of different types of high-affinity antibodies which can bind to different epitopes of the same antigen (polyclonal antibodies, pAbs), and resulting pAbs can be used as highly stable antitoxins or highly robust biosensors for food and environmental control.
However, if the purpose is the use of these antibodies in therapy, research (e.g. ELISA, Western Blot, Immunoprecipitation) or diagnostics, effector B cells need to be extracted from the spleen of immunized hosts and the most effective germlines need to be selected for further production of monoclonal antibodies (mAbs, specific to a single epitope of the antigen). The development of therapeutic antibodies, should also take into consideration the homology of the antibody to the patient’s immune system. For this reason, therapeutic antibodies generated in vivo in non-human hosts require the use of specific methodologies:
- Antibody generation in transgenic mice with human immunoglobulin genes
- Antibody generation in animal hosts, followed by antibody humanization

Hybridoma generation or phage display? Which method is the most adapted to your project?
Discover the answerIn vitro custom antibody generation
Many different in vitro development techniques are in use today. These techniques rely on the use of naïve, immune, or mutated/synthetic libraries containing the genetic information of different regions of antibodies (Fab, scFv, VHH) from different hosts (human, mice, rabbit, alpaca, among others).
The use of these techniques reduces the use of animals for antibody generation, instead, they rely on different in vitro systems, namely:
- Phage display
- Yeast display
- Ribosome display
Phage and yeast vectors express the variable domain of the antibody on their surfaces and, for this reason, can quickly be screened and selected according to their affinity to the target antigen. For instance, phage vectors can easily be screened and selected by biopanning, a technique that results in the enrichment of the phage particles displaying the variable domains with highest affinity to the target antigen. While yeast presenting exogenous antibody fragments can be screened and sorted by flow cytometry using a fluorophore-labeled epitope.
Contrarily, ribosome display is a cell-free technology used for in vitro antibody generation. Since this technique is not limited by time-consuming cell transformation steps, it can be used to screen much larger libraries than the ones currently possible using phage or yeast display technologies. However, this technique is also much more costly and technically demanding in comparison to conventional display systems. Ribosome display still presents some important technical limitations, including the reduced stability of the mRNA-ribosome-polypeptide ternary complexes and mRNA molecules, which are highly susceptible to degradation by ubiquitous and hard to eliminate nucleases.
These technologies have considerable advantages in comparison to in vivo development. For instance, since this process bypasses the immune system of the host, these antibodies can be designed to bind toxic or non-immunogenic molecules. Furthermore, they are also compatible with high-throughput screening procedures, which have the potential of improving this process in terms of parallelization, automation, and miniaturization.
Interestingly, the antibodies obtained by in vitro development and maturation require fewer quantities of the target antigen than immunization-dependent processes. This occurs due to the limitation of the B cell activation process, which requires at least 100 pM of a given antigen to take place.
Furthermore, these techniques allow a degree of control and selection that cannot be reproduced by in vivo development of antibodies. For instance, since screening conditions can be thoroughly defined, it is possible to generate antibodies that bind to a specific epitope of an antigen or to a specific antigen conformation, distinguishing between different post-translationally modifications of the same antigen. At the same time, antibody affinities can also be easily broadened or narrowed according to the objectives of each particular project.
Concluding remarks
Today, the choice of a proper antibody generation method depends on the goals of each specific project. For instance, in vivo methods generate antibodies with high affinity to the target antigen in a cost-effective way. These methods are ideal to generate antibodies intended for analytical purposes (e.g. diagnostics, environmental and food monitoring).
However, if the antibodies are intended for therapy and if they are generated in non-human hosts, it is necessary to introduce extra steps in the process to engineer the antibody and increase its compatibility with the human immune system (e.g. antibody humanization, use of transgenic animals with human immunoglobulin genes).
Moreover, in vitro custom antibody generation methodologies can be accomplished in much shorter timeframes and are compatible with high-throughput screening and molecular engineering techniques that allow a much quicker affinity maturation process. For this reason, they are generally recommended for the generation and development of therapeutic antibodies or to generate antibodies able to bind non-immunogenic molecules.
- Bradbury, A. R. et al. Beyond natural antibodies: the power of in vitro display technologies. Nat Biotechnol. 2011. 29(3):245–254. doi:10.1038/nbt.1791
- Hurrell, J. G. R. Monoclonal Hybridoma Antibodies: Techniques and Applications. CRC Press. Boca Raton, Florida, USA. 1st Edition. 2017. 239 pp. ISBN: 9781315895727
- Li, R. et al. Ribosome display: a potent display technology used for selecting and evolving specific binders with desired properties. Mol Biotechnol. 2019; 61(1):60-71. doi: 10.1007/s12033-018-0133-0
- Persson, H. et al. In Vitro Evolution of Antibodies Inspired by In Vivo Evolution. Front Immunol. 2018. 9:1391. doi:10.3389/fimmu.2018.01391