Antibody production
Antibody production
How to choose the right antibody production process for your project
The value of the antibody-antigen interaction, recognized by the end of the 19th century, has given rise to the development of a wide array of immunotechnologies for both therapeutic and non-therapeutic applications. Presently, many antibody production processes are available for large-scale applications, for this reason, it is crucial to understand the advantages and limitations of each process to choose the best fit for each specific application.
Antibody applications
Antibodies are highly sensitive and specific biomolecules, with the ability to detect low quantities of antigens in samples including drugs, toxins, small proteins, virus, and even pathogens. Antibodies are recurrently used for biomedical research (ELISA, Western Blot, Immunohistochemistry, Immunofluorescence Microscopy, Immunoprecipitation, among others) and as highly sensitive biosensors for food safety and environmental monitoring applications.
Unlike expensive chromatographical and analytical techniques, the development of antibodies for monitoring purposes possesses the advantage of fast detection and sensitivity, while at the same time allowing the visualization and quantification of specific markers in environmental and clinical samples.
However, being a technically demanding process, the production of antibodies requires an in-depth knowledge of the available processes to choose the best one for each project and application.
In this article, we describe the main antibody production processes, as well as their advantages and disadvantages, to help you choose the one that best matches your unique requirements. You can also take our online quiz to help you decide the best solution according to the timeframe and funding of your project.
Antibody production processes
- Hybridoma technology
- Polyclonal antibody production
- Phage display technology
Hybridoma technology
Hybridoma technology was first developed by Milstein and Köhler in 1975. These two researchers discovered how to create a stable hybrid cell line able to secrete a high amount of antibodies specific to a unique epitope of an antigen, commonly known as monoclonal antibodies (mAbs).
The technique consists of extracting effector B cells from the spleen of previously immunized animals. These cell lines will only be able to replicate a reduced number of times in laboratory conditions before they die. For this reason, Milstein and Köhler fused these effector cells with immortalized myeloma cells to produce immortal effector B cells with the ability to secrete high quantities of mAbs, which were later named hybridomas.
The fusion of the two cell lines and subsequent selection of hybridomas is possible since the myeloma cells used in this process lack hypoxanthine-guanine-phosphoribosyltransferase (HGPRT), an important enzyme of the nucleotide salvage pathway. For this reason, to select positive hybridoma cells, the mixture needs to be cultured in a selective medium where de novo nucleotide synthesis is inhibited. In this medium, myeloma cells are non-viable, while effector B cells will die quickly, and only the fused hybridoma cells will be able to survive.
The advantage of this technique over others is its robustness, reproducibility, and the possibility to adapt for large-scale production. The limitation of this technique is that it will only produce fully animal antibodies (also known as murine antibodies if mice are used as the source of effector B cells). This is undesirable from a therapeutic point of view since animal antibodies can trigger the immune response in human patients resulting in reduced therapeutic effectiveness. Therefore, for therapeutic applications, these antibodies need to be humanized.
Applications: Suitable for projects that require large-scale production of stable antibodies with low risk of batch-to-batch variations. This technique provides the ideal solution for diagnosis applications.
Polyclonal antibody production
The production of polyclonal antibodies (pAbs) starts by triggering an immune response in an animal host (rabbit, chicken, mouse, rat, goat, sheep, llama, or alpaca). An effective generation of pAbs requires a careful design of the antigen and antigen-carrier protein, as well as the planning of an effective animal immunization strategy.
The initial immune response against a low amount of the antigen will trigger the development of different germlines of effector B cells and memory B cells producing antibodies able to bind different epitopes of the same antigen. While subsequent immunizations with the same antigen ensure the activation of the several memory B cells germlines and successful in vivo affinity maturation of the pAbs.
Antibody expression levels are measured throughout the immunization process to ensure specificity and adapt subsequent antigen dosages. When desirable levels of antibody production are achieved, the blood of the immunized animal is harvested and the cellular and serum fractions are subsequently separated. The serum of the animal containing high-affinity and high-specific pAbs (also called antiserum), can be further purified against the specific antigen.
An advantage of pAbs in comparison to mAbs is that they can bind to multiple epitopes of a specific antigen, ensuring that even if a specific epitope of the antigen suffers modification, these pAbs will still be able to recognize the molecule. This technique can also generate high amounts of pAbs, and it is also fast and inexpensive in comparison to the hybridoma technology. These antibodies are also more robust than normal mAbs, and they remain stable under a wide range of environmental conditions (pH, salinity, temperature). However, the downside of this technique’s versatility is its reduced reproducibility, which leads to extensive batch-to-batch variations.
Applications: The antiserum of immunized individuals is still the only effective way to treat diseases like Ebola, which act too quickly in the organism to allow the development of an effective adaptive immune response. They are also recurrently used as antitoxins, to treat patients that have been bitten or stung by venomous and poisonous animals. pAbs have limited use in diagnosis due to the high possibility of false-positives. However, due to their stability, these antibodies are recurrently applied for detection and monitoring purposes in environmental and clinical settings.
Phage display technology
Phage display, first demonstrated by George Smith in 1935, has been recurrently used for the production of recombinant monoclonal antibodies (rAbs). This technology allows the development of antibodies from different sources through the clonal selection of antibody fragments in a bacteriophage coupled with a prokaryotic system (usually Escherichia coli).
Antibodies obtained through this technique derive from antibody libraries containing genomic information of the variable domains. These libraries can either be naïve or immune libraries, depending on if they were obtained from non-immunized or immunized hosts (human, mouse, rabbit, alpaca, among others), respectively. The latter is a more time-consuming and expensive process, requiring the construction of a unique library for each specific antigen. However, since these libraries originate from immunized hosts, they are already highly enriched for antibodies with high affinity towards the target antigen. Moreover, through random and site-directed mutagenesis, these libraries can be used to improve the affinity of specific antibodies and improve their biophysical characteristics.
Contrarily, naïve libraries are cheaper, easier to obtain, and usually used for the development of antibodies against an unlimited number of different antigen structures. However, these libraries need to be several degrees of magnitude more diverse than immunized libraries, to increase the chances of obtaining antibodies able to bind a wide variety of antigen structures.
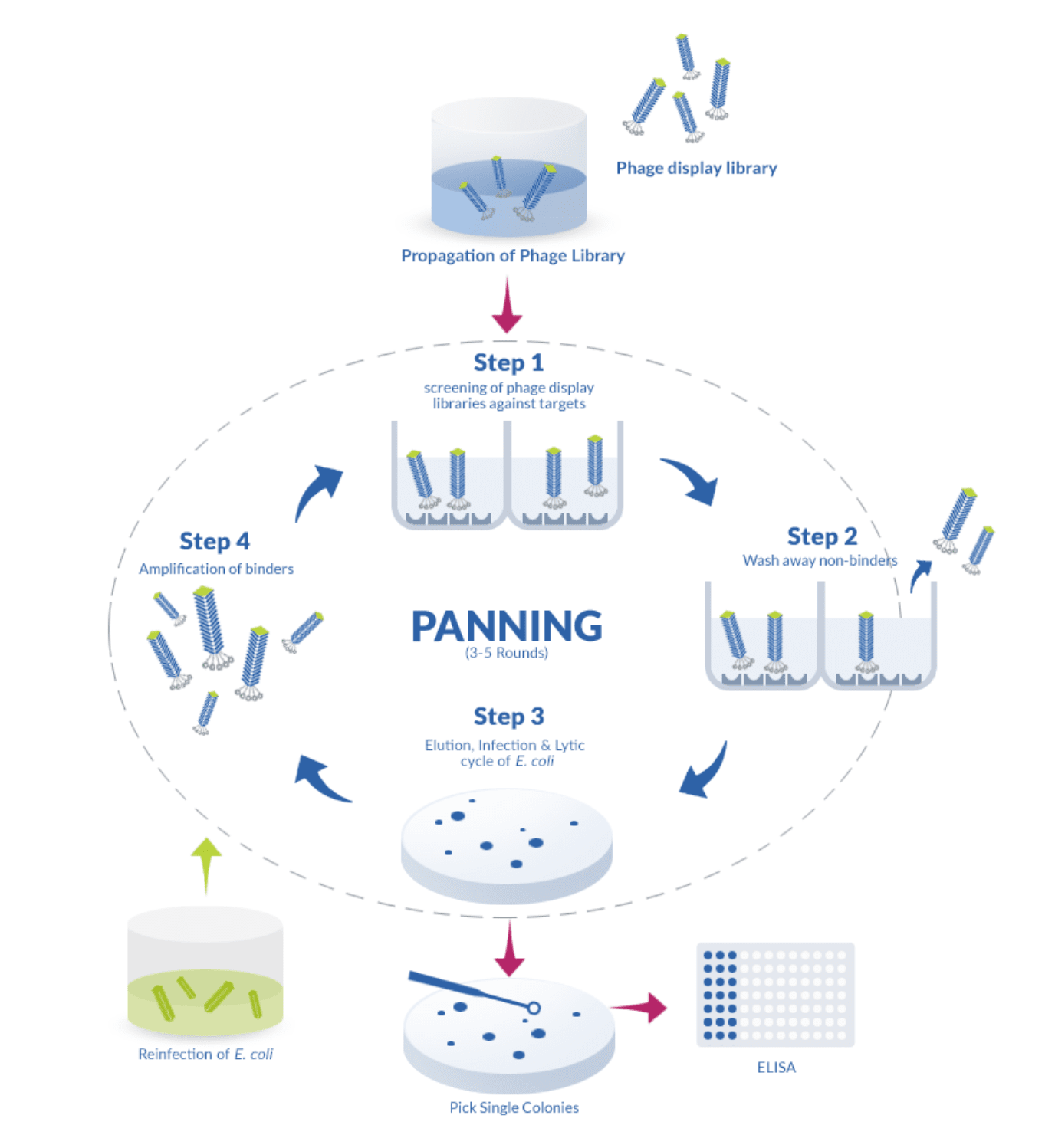
The process for the construction of these libraries is simple since it relies on the reverse transcription of mRNA followed by PCR amplification with specific primers designed to amplify the heavy (H) and light (L) chains of the variable domain. These genes are later fused with the coat proteins of a phage (typically the filamentous phage M13), which then becomes able to display the binding-domain of the antibody on its surface. For this reason, phages displaying a wide variety of binding-domains can quickly be screened against a particular antigen, simplifying the choice of a suitable phage, which can afterward be propagated on its prokaryotic host.
The main advantage of this technique is that it is fast and it does not require the immunization of animals and, at the same time, it allows the development of fully human or fully animal (mouse, sheep, llama, rabbit) antibodies. In this way, for therapeutic applications, it poses a significant advantage in comparison to the hybridoma technology since it eliminates the need for the time-consuming process of antibody humanization. Moreover, since antibody development occurs in vitro, it bypasses the limitations of the immune system by allowing the development of antibodies able to bind very small or non-immunogenic molecules.
This technique also requires the use of less purified antigens in comparison to the hybridoma and polyclonal antibody technology, in which the antigens need to be carefully designed, purified and conjugated to trigger an adequate immune response.
Applications: Antibodies developed through the phage display technology are preferentially used for therapy and diagnostics. This technology represents the “gold standard” for therapeutic antibody development as it allows to circumvent the time-consuming antibody humanization process. It is also the technology of choice to develop antibodies against non-immunogenic molecules.
- Amine, A., Mohammadi, H., Bourais, I., Palleschi, G. Enzyme inhibition-based biosensors for food safety and environmental monitoring. Biosens Bioelectron. 2006; 21(8):1405-1423. doi:10.1016/j.bios.2005.07.012.
- Bazan, J., Całkosiński, I., Gamian, A. Phage display—A powerful technique for immunotherapy. 1. Introduction and potential of therapeutic applications. Hum Vaccin Immunother. 2012; 8(12):1817–1828. doi:10.4161/hv.21703.
- Carvalho, L. S., da Silva, O. B., de Almeida, G. C., de Oliveira, J. D., Parachin, N. S., Carmo, T. S. Production Processes for Monoclonal Antibodies. Fermentation Processes. 2017. doi:10.5772/64263.
- Köhler, G., Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature. 1975; 256:495–497. doi:10.1038/321522a0.
- McCafferty, J., Griffiths, A. D., Winter, G., Chiswell, D. J. Phage antibodies: filamentous phage displaying antibody variable domains. Nature. 1990; 348(6301):552-554. doi:10.1038/348552a0.
- Smith, G. P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science. 1985; 228(4705):1315-1317. doi:10.1126/science.4001944.