 Peptide synthesis
Peptide synthesis
Are peptide drugs still in use for therapeutic applications?
Peptide drugs have been in use for almost a century. Initially, these molecules served only as hormone analogs to balance metabolic diseases. Today, peptides can be easily synthesized and modified chemically to achieve many complex tasks within our organism including crossing membranes and reaching intracellular targets. Decades of peptide development have led to the discovery of cell-penetrating peptides and cyclic peptides, both with promising results for different therapeutic applications.
Discovery and limitations of peptide drugs
Peptides are regulatory molecules that mimic natural hormones. The first of these hormone analogs ever to get approval for clinical use was insulin.
Since the introduction of this hormone-based therapy in the 1920s, more than 60 peptide drugs have been approved for clinics in the United States, Europe, and Japan. And every year, about 20 new peptides enter into clinical trials.
Peptide drug sales exceeded 70 billion USD in 2019 alone, and the market is expected to continue to grow at 9.1% per year at least for the next 5 years.
The success of these molecules relies on the essential role they play in our organism. Peptides are natural signaling molecules produced across all domains of life. Beyond their role as hormone analogs, peptides can also be used to interfere with normal protein-protein interactions. In this not surprising though that the vast majority of peptide drugs approved for therapy are predominantly used to treat metabolic disorders and cancer.
Protein-protein interactions are a crucial and fundamental stage of many essential biological processes. Several processes including metabolism, biosynthesis, transport, transcription, translation, and post-translational modifications depend on the activity of natural receptors. Peptide drugs regulate these processes by either activating (agonists) or blocking (antagonists) specific cell receptors and, thus, enhance or disrupt specific cellular processes.
Thus, the clinical success of these drugs depends on the identification of the best target and optimization of the peptide’s binding efficiency. These steps pose important challenges for computational biologists and biochemists.
Nevertheless, peptides are much more effective than small molecules at disrupting protein-protein interaction. Thus, they are often referred to as interfering peptides.
Overcoming the natural limitations of peptides
Peptide drugs have three major limitations that hinder their therapeutic effectiveness: in vivo instability, poor oral adsorption, and membrane impermeability. Their instability originates from the quick degradation of peptides on our organism by widespread proteolytic enzymes. For this reason, they have poor oral absorption, therapeutic peptides should be mainly administered by injection.
This instability is also regarded as an advantage by researchers developing peptide drugs. Their inherent instability makes them less toxic than other small synthetic molecules. Moreover, they are not prone to accumulating in the organism, unlike other small synthetic drugs.
To decrease their susceptibility to proteolysis, peptide drugs have been routinely modified by chemical processes to increase their robustness. These processes comprise:
- Termini protection – peptidases including serum aminopeptidases and carboxypeptidases, can degrade peptides from their N- and C-termini. These terminal amino acids may be replaced by residues that are naturally more resistant to degradation (e.g. methionine, serine, alanine, valine, among others), by unnatural amino acid analogs or by functional groups (C-terminal amidation or N-terminal acetylation).
- Replacement by enantiomers on the peptide backbone – in living systems, all amino acid residues are in the form of the L-enantiomer. The replacement of residues within the peptide by D-enantiomer may confer further resistance to proteases, as these enzymes cannot recognize nor cleave the unnatural enantiomer. However, this modification needs to be carefully considered as it can lead to important conformational changes and compromise the in vivo activity of the drug.
- Peptide cyclization – cyclized peptides can be synthesized by linking the terminal regions or side chains of these drugs with amide, lactone, ether, thioether or other types of chemically stable bonds. Cyclic peptides are known to be more resistant to proteolytic degradation due to the lack of both amino and carboxyl termini. They also possess an enhanced biological activity and even to have an increased membrane permeability in comparison to their linear counterparts, although the exact mechanisms are not fully understood.
- Peptide conjugation – peptides can be conjugated with molecules that are known to naturally cross membranes. These molecules, named cell-penetrating peptides, are known to enhance the active transport of peptide drugs. However, the conjugation may also increase the immunogenicity of these drugs, thus their ability to cause an immune response should be carefully analyzed.
New approaches for peptide drug discovery
Many peptide drugs currently in use have originated from natural molecules. But the inherent instability of natural peptides has led to the development of different approaches to enhance peptide discovery.
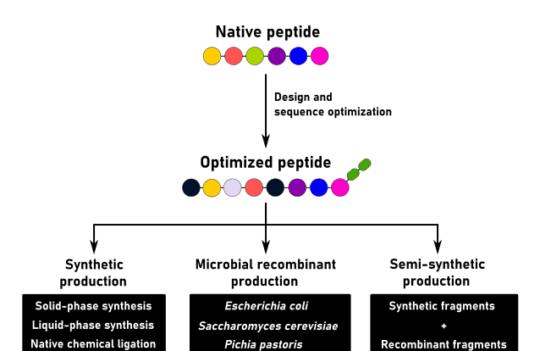
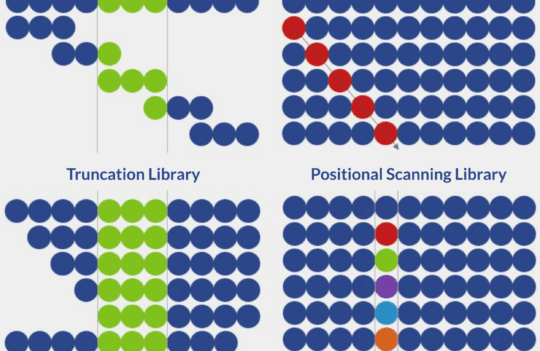
While new peptide drugs can be generated by chemical modification of natural peptides, there is an increasing interest in the development of high throughput approaches. Two of the most popular approach for peptide discovery are genetic and synthetic approaches.
Genetic approaches consist of the construction and screening of DNA libraries by common display methods. Today, phage display is the most used method to identify peptides with the highest affinity to specific protein receptors. Phage particles displaying peptides on their surfaces can be enriched by biopanning and the highest affinity peptides can be further recombinantly produced by high-performing microbial systems.
An obvious advantage of this method is that peptide sequencing becomes unnecessary, as the DNA sequences are already available. However, since these peptides are produced by a recombinant organism, the resulting drugs cannot contain unnatural residues or chemical groups.
New peptide drugs can also be discovered using synthetic approaches. However, this approach requires proficiency in the use of complex computational biology tools and a solid synthetic peptide database. Besides, to use this in silico approach, the structure of the target protein needs to be known.
An advantage of these methods over genetic methods is their ability to produce modified peptide drugs. Moreover, these drugs can be produced in a cost-effective way using chemical processes such as solid-phase peptide synthesis.
Concluding remarks
Peptide drugs have been in use for the treatment of metabolic diseases for almost a century. These drugs present several advantages in comparison to small chemical molecules. They are less toxic and more specific than most of these drugs, however, they are also very unstable and naturally unable to cross membranes.
The limitations of natural peptides have prompted the development of new peptide drugs. In this context, cell-penetrating peptides and cyclic peptides comprise the most promising breakthroughs obtained in the past decades.
These peptides offer interesting solutions to one of the greatest challenges of modern medicine: the ability to cross membranes and reach intracellular targets. In this way, it is estimated that peptide development and discovery will continue to interest members of the scientific and industrial community.
- Joo, S. H. Cyclic Peptides as Therapeutic Agents and Biochemical Tools. Biomol Ther (Seoul). 2012; 20(1): 19–26. doi: 10.4062/biomolther.2012.20.1.019
- Lau, J. L. and Dunn, M. K. Therapeutic peptides: Historical perspectives, current development trends, and future directions. Bioorg. Med. Chem. 2018; 26(10):2700-2707. doi: 10.1016/j.bmc.2017.06.052
- Lee, A. C. et al. A Comprehensive Review on Current Advances in Peptide Drug Development and Design. Int J Mol Sci. 2019; 20(10):2383. doi: 10.3390/ijms20102383
- Molek, P. et al. Peptide phage display as a tool for drug discovery: targeting membrane receptors. Molecules. 2011; 16(1):857-887. doi: 10.3390/molecules16010857