 Antibody production
Antibody production
When you should choose to develop a peptide antibody for your project
The discovery of peptide antibodies has reduced antibody generation costs and increased the specificity of natural monoclonal antibodies. These antibodies remain highly useful for the generation of therapeutic and diagnostic antibodies as they allow greater control over the target-specificity of the final products. Check out our frequently asked questions (FAQs) page about monoclonal antibodies on our dedicated page.
Peptide antibodies: what are they and what are they used for?
Peptide antibodies were first described in the 1980s, soon after the discovery of the hybridoma technology by Köhler and Milstein. The intended use of these specific antibodies was to direct the antigen-antibody recognition sites to specific regions (i.e. peptides) of the native protein.
They became popular due to the relatively simple and inexpensive peptide production processes and due to their potential to enhance the specificity of monoclonal antibodies.
The use of peptides as immunogens allowed researchers to control which epitope, or epitopes, of an antigen were recognized and targeted by the host immune system during the antibody generation stage. Later, the use of these peptides also allowed the cost-effective design of antibody library screening techniques.
Peptide design and selection continues to evolve and it increasingly takes into account the properties of each peptide region and their potential for immunogenicity and antigenicity. Commonly, peptide immunogens are designed to mimic:
- Terminal or internal specific sequences of the native protein
- Specific proteolytic degradation products
- Specific posttranslational modifications
- Phosphorylation, sulfation or citrullination sites, among others
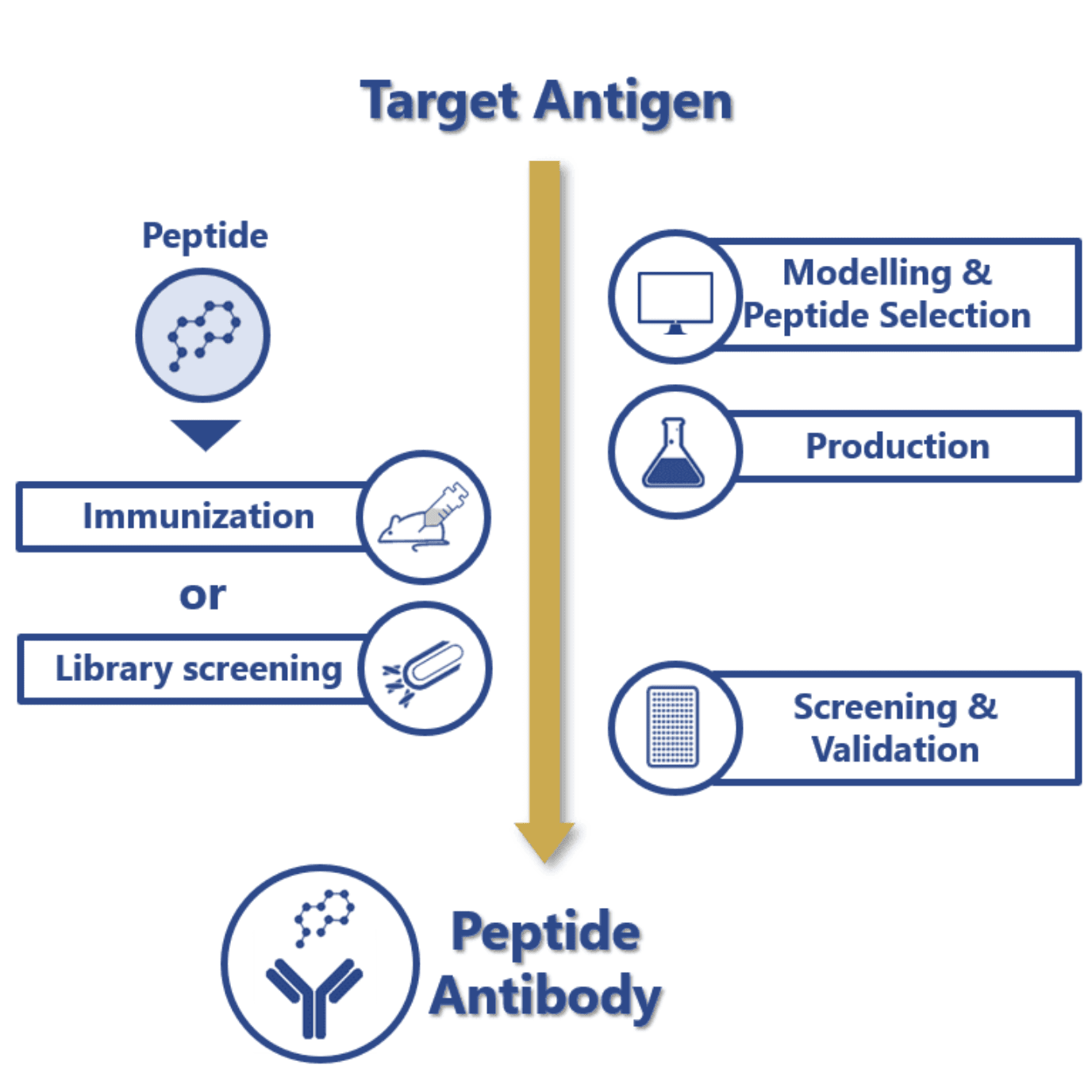
Development of peptide antibodies
Animal immunization or library screening
Immunization is the key initial stage of many in vivo and in vitro antibody generation processes. For this reason, immunogen selection and design continue to be the most crucial phases of this process.
Traditionally, proteins have been used as effective immunogens for antibody generation. But there are some drawbacks in using native proteins for this purpose. For instance, in some cases the native protein is unstable, hard or costly to produce. Moreover, when using the entire antigen for immunization it is not possible to control which epitope of the antigen will be recognized and bound by antibodies generated in vivo.
On the contrary, peptide immunization allows to exert a greater control on the specificity and affinity of in vivo generated antibodies, because researchers may choose to target a specific region(s) of the antigen. The chemical synthesis of defined epitopes is easy to perform and inexpensive. Peptides are also known to be more stable and soluble than native proteins. Moreover, the use of peptides for immunization also reduces the risk of cross-reactivity with other non-specific antigens.
More recently, it is becoming increasingly popular to develop mutation-specific antibodies. These antibodies are designed only to recognize the mutated target and not the native protein. For this reason, they are frequently designed to target tumor markers.
Custom antibody generation using in vitro display techniques, such as phage display, has also profited from the inclusion of peptides in the screening stage. For instance, the use of peptides instead of the full-length native protein allows to decrease the cost of antibody-antigen high-throughput screening processes. Moreover, similarly to in vivo antibody generation processes, the use of peptides allows the recovery of epitope-specific antibodies and, for this reason, the selection of highly specific-antibodies.
Parameters to consider during peptide design
Peptides used for immunization should present two distinctive characteristics, antigenicity and immunogenicity, while peptides used for antibody library screening approaches only need to have antigenicity.
Antigenicity is the ability of a peptide to mimic the antigen-antibody binding site. This will ensure that the peptide antibody will be able to recognize the native protein. This characteristic is easier to control during the planning and design stage and usually about 50% of the designed antibodies present the proper affinity to the intended target.
Whereas immunogenicity is the ability of a peptide to challenge the immune system of the host. This is more difficult to control and to predict. And to increase their immunogenicity, these peptides should be long enough to activate T and B cells, or else, they should be conjugated with a carrier protein able to trigger the immune response by itself.
The choice of the proper peptide target is crucial and must be carefully considered taking into account the specific downstream application of the peptide antibody. For this reason, peptide antibodies targeting denatured protein targets are much easier to design than antibodies targeting antigens in their natural conformation. This happens because most regions of the denatured protein are easily accessible, while many regions of the native proteins may be inaccessible to the large monoclonal antibodies.
In these cases, the specificity of the peptide antibody should be confirmed and validated, when possible, against the complete target antigen in the same structural conformation as the one that will be used in the final assay. This is an important step because solid-surface immobilized antigens may present a different structural conformation when compared to the same antibody in liquid preparations.
Parameters such as hydrophilicity, flexibility, surface exposure and charge distribution should be included in the early steps of peptide selection. This requires that the structure of the native antigen is known either by experimental evidence or by protein modeling approaches.
| Parameter | Recommended | Additional considerations |
|---|---|---|
| Peptide size | Ideally between 8-20 amino acids
Recommended: 15 amino acids |
Peptides with this length range are preferable in order to minimize solubility problems and to decrease the risk of secondary structure formation that may be too different from the native antigen structure.
When using peptides ≤ 15 aa it is usually advisable to use peptide-carrier conjugates or multiple antigen peptide systems to ensure their immunogenicity. |
| Amino acid composition | Hydrophobic amino acid content ≤ 50%.
Sequences without multiple and adjacent residues (Val, Ile, Tyr, Phe, Trp, Leu, Gln, Thr). Low amount of Gln, as these residues tend to form extensive hydrogen bonds in the peptide structure. Presence of Tyr and Pro residues tends to stabilize the peptide structure. |
Peptides rich in hydrophilic and charged amino acids are preferred for specific natural epitope targeting because they are usually located on the surface of the antigen.
However, if the purpose of the project is to detect epitopes hidden in the native protein structure (cryptopes, that become available after denaturation) is it better to enrich with hydrophobic amino acids instead. |
| Peptide structure | Linear and flexible.
Or containing a defined secondary structure conformation (cyclic, with loops and turns) mimicking the native protein. |
Although controversial, some researchers have suggested that the presence of secondary structure is often desirable and known to produce peptide antibodies with high affinity.
It is also known that the use of linear peptides generates antibodies able to target linear epitopes or denatured antigens. |
| Protein target | Peptides can be designed to target highly variable or highly conserved protein regions.
Peptide antibodies targeting terminal regions are more likely to recognize the native protein. |
Peptide antibodies intended for a broad recognition range should be designed to target conserved domains. Whereas, peptide antibodies targeting a specific protein should target hypervariable domains. |
Concluding remarks
The development of peptide antibodies has allowed researchers a greater control and flexibility in antibody generation using in vivo or in vitro techniques.
However, since peptides represent and target only specific and small regions of the target protein, several parameters should be taken into account during the planning and design of these peptide immunogens.
The choice of an adequate peptide immunogen may lower antibody production costs and also increase the specificity of natural antibodies. For this reason, the use of these antibodies can be an advantageous approach for therapeutic and diagnostic applications.
- Trier, N. H. et al. Production and characterization of peptide antibodies. Methods. 2012; 56(2):136-44. doi: 10.1016/j.ymeth.2011.12.001
- Trier, N. H. and Houen, G. Peptide Antibodies in Clinical Laboratory Diagnostics. Adv Clin Chem. 2017; 81:43-96. doi: 10.1016/bs.acc.2017.01.002
- Lee, B. S. et al. Production of antipeptide antibodies. Methods Mol Biol. 2010; 657:93-108. doi: 10.1007/978-1-60761-783-9_7
- Lee, B. S. et al. Antibody Production with Synthetic Peptides. Methods Mol Biol. 2016; 1474:25-47. doi: 10.1007/978-1-4939-6352-2_2